Recently Published
Candidate gene search tutorial
Some methods to look for candidate genes using qtlTools

sim.ct
Simulation of cis-trans test performance
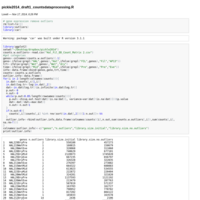
ph2015_snpsfromase_tojoinmap.R
Workflow from ASE counts to joinmap input file

maizeqtlmapping.R
Basic, 1-way QTL mapping for Maize P experiment
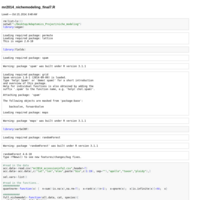
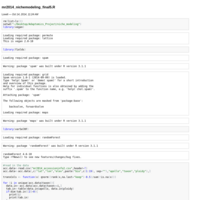
mr2014_nichemodeling.R
Intro for niche modeling script. For Martin
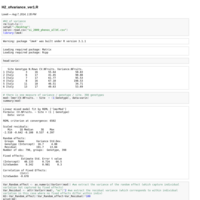
H2_ofvariance_ver1.R
First try at heritability of variance
dd2014_qtlmappingstatisticsplotting_draft3.R
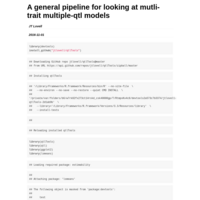
full qtl, gene expression and covariate analysis

dd2014_qtlmappingstatisticsplotting_draft2.R
Basic QTL analysis without epistasis. Generates a table of statistics for each QTL*trait combination

dd2014_qtlgeneexpression_draft2.R
This script conducts simple statistics looking for differential gene expression between alleles along a QTL interval. This provides a way to narrow down the number of potential genes underlying a QTL by comparing the patterns of gene expression with phenotypic responses to the different environments

dd2014_qtlmappingstatisticsplotting_draft1.R
Full QTL analysis in one script, all automated
rl2014_allqgen_ssr_analyses.R
All analyses of qgen and ssr gen so far.
Basically plotting everything.

Adaptomics_CarbonIsotopeAnalysis_ComparisonofExperiments
We did two separate drydowns because there was very poor germination of a few genotypes in our first try and we did not have enough seed to re-do the drydown for a few of the genotypes.
So, genotypes are nested within experiments... that is, all genotypes are represented in only 1 experiment. This is not an ideal design, especially since we see an "experiment" effect.
Lets discuss.
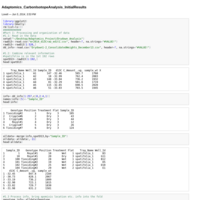
Adaptomics_CarbonIsotopeAnalysis_InitialResults
Some comparisons of water-use-efficiency (d13C), treatments (wet/dry), populations and mating systems

rl2014_allnewanalyses.R
New analyses of molecular and quantitative genetic diversity showing all data (without means)