Recently Published
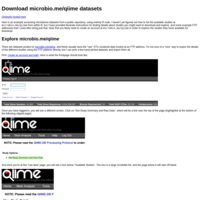
Download microbio.me/qiime datasets
Download publicly available datasets from microbio.me/qiime directly from within R using phyloseq and a little function I provide.

biom: An interface package (beta) for the BIOM file format
This is an R package for interfacing with the BIOM format. Currently in beta form, this package includes basic tools for reading biom-format files, accessing and subsetting data tables from a biom object (which is more complex than a single table), as well as limited support for writing a biom-object back to a biom-format file. The design of this API is intended to match the python API and other tools included with the biom-format project, but with a decidedly “R flavor” that should be familiar to R users. This includes S4 classes and methods, as well as extensions of common core functions/methods.

plot_heatmap
A special implementation of heatmap plots, inspired by the NeatMap package and based exclusively on ggplot2 graphics, implemented and freely available in the phyloseq package

Some bacterial genome analysis using R
Preliminary examples for a single class sectional displaying "Finding Patterns" in bacterial genome analysis

Re-analyzing the enterotype dataset - part 1
This part notes the omission of samples from the Sanger data that, if included, change their result of 3 clusters

Test rmd in RStudio with simple example using phyloseq package
An example using the phyloseq package to plot a network representation of species (cophenetic) distances.


